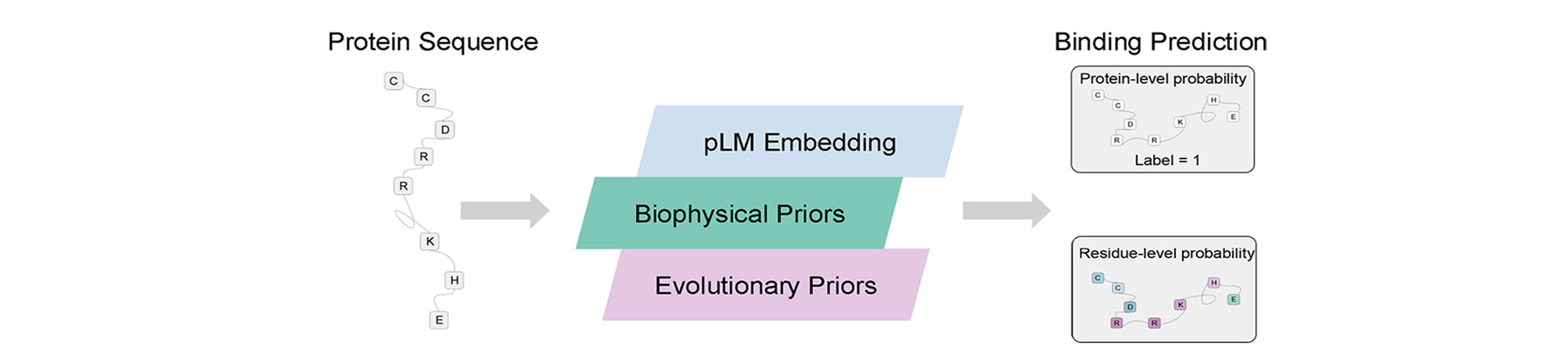
ABSTRACT: Protein interactions with nucleic acids are fundamental to numerous biological processes. Here, we present PNABPred, a multi-modal framework that integrates biophysical and evolutionary priors into a protein language model to predict protein-nucleic acid interactions. Specifically, semantic representations are combined with classical evolutionary representations and biophysical representations. Our results showed that PNABPred outperforms other state-of-the-art sequence-based methods. In RNA-binding protein classification tasks, PNABPred achieved an MCC of 0.889 and AUROC of 0.990, 17.44% and 3.23% higher than the next best method (Seq-RBPPred), respectively. In DNA-binding site prediction, PNABPred also outperformed the transformer-based method CLAPE-DB by 20.31% and 4.65% (MCC 0.468; AUROC 0.922) on the Test_129 dataset. PNABPred employs only protein sequences as inputs, identifying nucleic acid binding sites even in intrinsically disordered regions. This framework supports scalable sequence screening and annotation of nucleic acid binding proteins for basic research, biotechnology, and therapeutic development applications.
For detail: https://doi.org/10.1016/j.isci.2026.115795